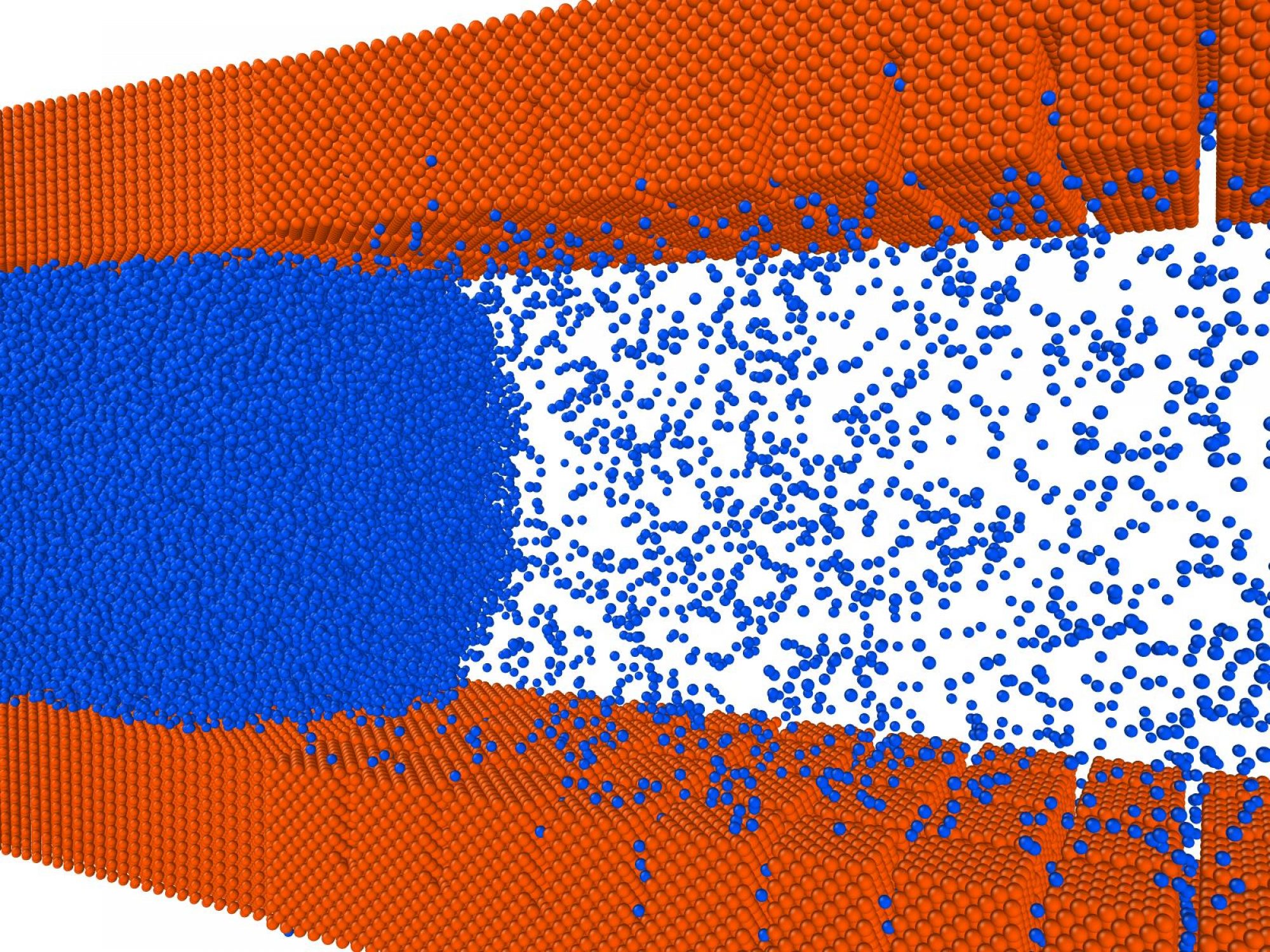
We develop and employ computational methods to predict molecular-scale features of systems that show the desired response to the process conditions. We are particularly interested in systems whose response can be depicted by the adjacent figure’s free energy cycle. Here, x denotes the physical state of the system, and lambda denotes the process condition. The cycle remains fundamental to our group’s scientific investigations, as seen from the following projects:
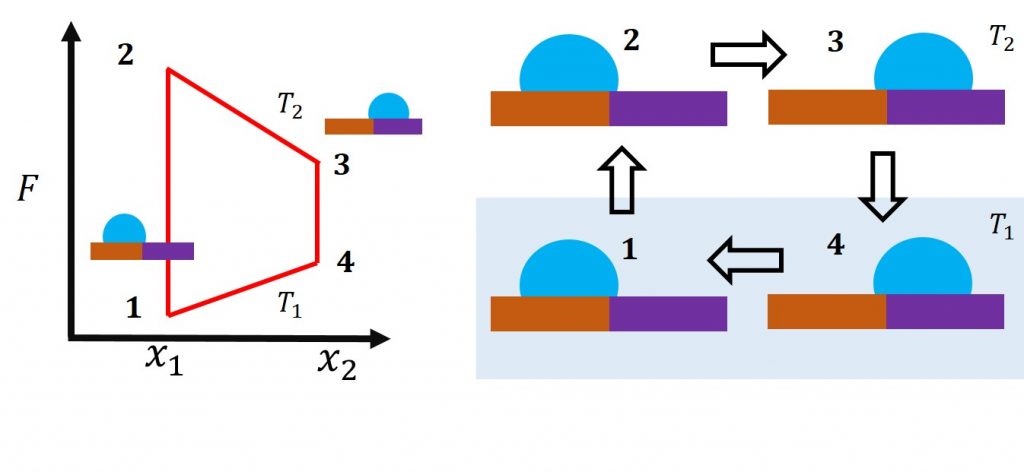
Droplet Engine: We are designing solid surfaces that result in a liquid droplet’s desired motion in response to the change in external conditions. We use the thermodynamics of solid-liquid and liquid-vapor interfaces, statistical mechanics, molecular simulations, and the principle of maximum entropy to predict the spatial variation of surface features. The droplet-solid system can be considered as an “engine” that performs the work of translating the liquid droplet along the surface. Its behavior can be depicted by the adjacent figure’s free energy cycle, where the external condition is temperature.
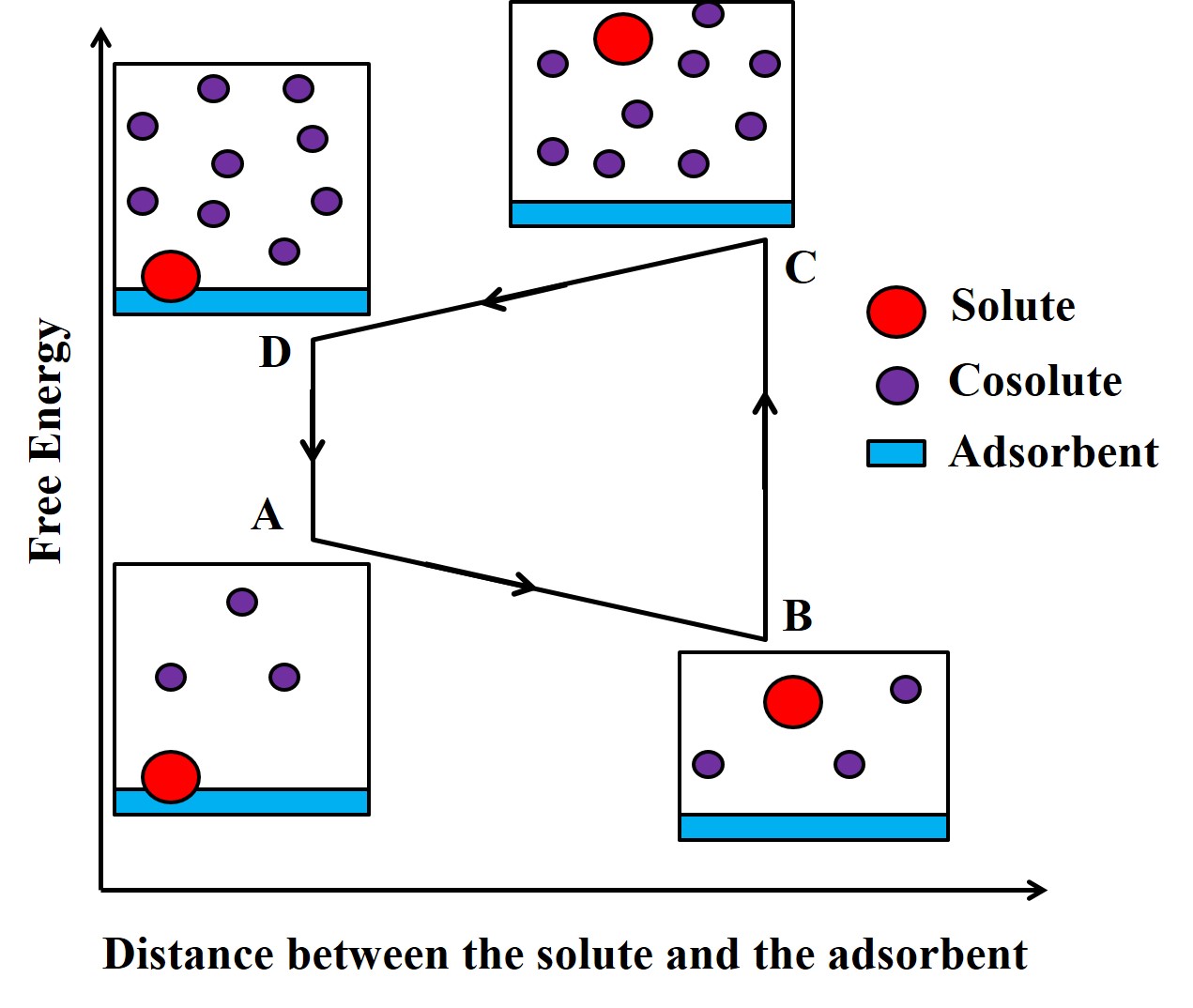
Smart adsorbents: We are designing adsorbents that adsorb specific solutes in response to the presence of specific cosolutes. Such “smart” adsorption behavior can help minimize expensive processes that are required for the regeneration of adsorbents. We use the thermodynamics of soft matter, statistical mechanics, molecular simulations and the principle of maximum entropy to predict the chemical nature of the above adsorbents. The process of smart adsorption can be represented by the adjacent figure’s free energy cycle.
Computational mutagenesis of biological motors: The biological cells are
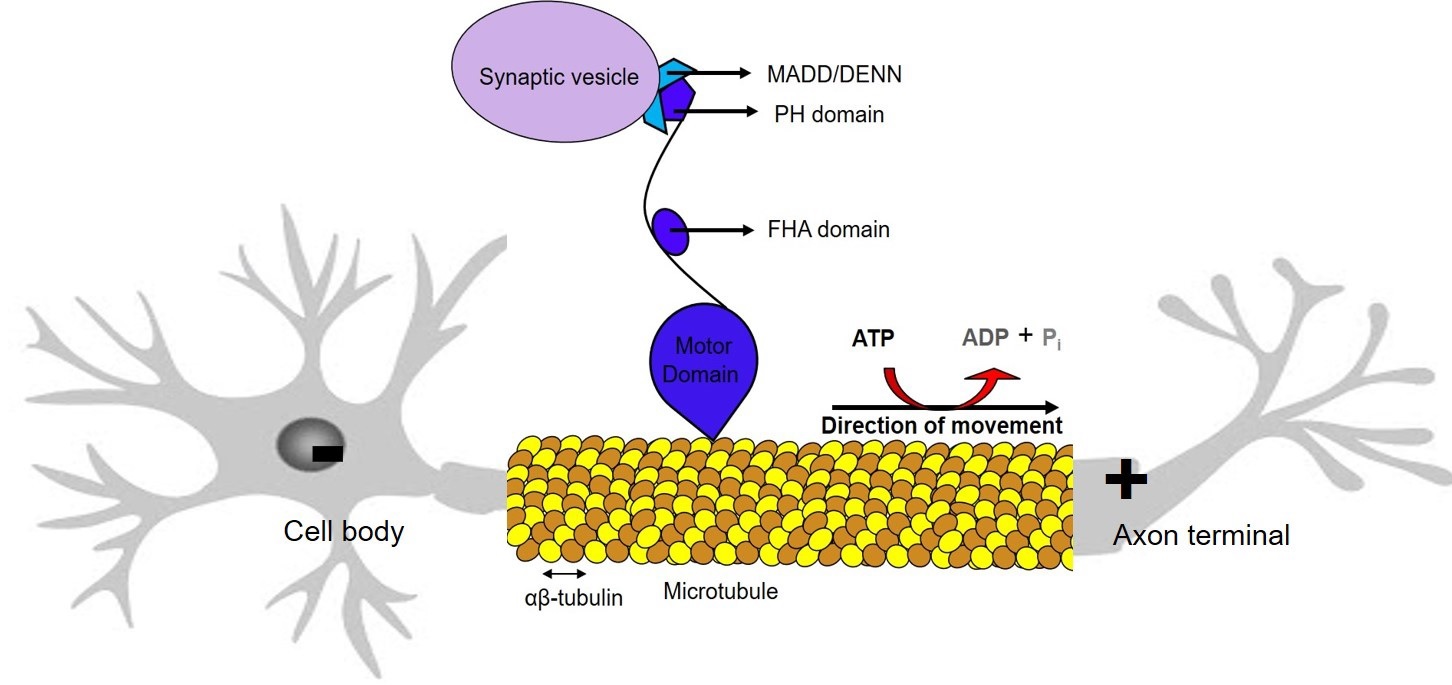
composed of several motors that produce useful work by using the chemical energy. These motors are made of proteins, and their operation depends on the chemical composition of the protein molecules. The mutagenesis studies aim to understand the effects of changing the above chemical composition. We are developing multiscale methods to perform these studies computationally. Though the behaviour of the motor-proteins is characterized by the complex free energy landscape, and the system is off-equilibrium, the computational strategies used to handle the thermodynamic cycle shown above are also applicable here.
